I’ve just released a new homepage with a succinct summary of what you can do at DNA Painter. In addition to this, I’m presenting this new page answering some of the questions that come up again and again.
If you’re completely new to the site, I recommend first reading the help page What is DNA Painter?
If you have a question that isn’t answered here or in the main FAQ, please feel free to email info@dnapainter.com. Thanks!
Chromosome mapping
Where do I upload my raw DNA file?
This is perhaps the top question, and has a simple answer: nowhere
- DNA Painter does not use raw DNA data at all!
I appreciate that this might be counterintuitive. After all, the idea of a system that can read in your DNA and automatically output a chromosome map showing which segments you inherited from your ancestors sounds amazing. And perhaps in the future, when tools are more advanced and databases are more integrated, this might be possible. But right now, this is not how chromosome mapping works.
Don’t I need my raw DNA to make a chromosome map?

No. When you click the ‘Create a new map’ button, DNA Painter will output a page with a virtual set of chromosomes. These represent you. They don’t contain any DNA data (the A/C/T/G values for each tested gene).
You’re able to map your chromosomes by combining genealogy with the DNA match information:
- Genealogy: your knowledge of how you are related to a match and the identity of the most recent common ancestors
- DNA match information: the chromosome, start and end point of each matching segment (also referred to as the segment data), which you get from FamilyTreeDNA, Gedmatch, Living DNA or MyHeritage.
For more information, read the ‘how to use’ help page or this step-by-step blog post.
Can I use matches from AncestryDNA for chromosome mapping?
No. In order to map the segments you share with matches, you need the coordinates (chromosome number, start and end position) of each segment you share with them. AncestryDNA does not provide this data, although you can ask your matches to upload their DNA to a site that does. AncestryDNA provides the amount of shared DNA, which you can use in the Shared cM Project and the What are the Odds? tools. To find other tools you can use with AncestryDNA information, visit the tools page and filter the list using the AncestryDNA checkbox.
What is the point of mapping your chromosomes anyway? What does it tell you?
For more information on this process, along with a discussion of how it can (and can’t) help you, please read Why map your chromosomes?
I just don’t get it. Who can explain it to me?
It’s probably less complicated than you think. You don’t need any deep scientific knowledge. Before you start, you just need to understand:
- You have a set of chromosomes from your father and another set from your mother
- This means you have two copies of each numbered chromosome: for example, one paternal copy and one maternal copy of chromosome 1
Your DNA testing company can tell you which chromosomes you match on and the position of each segment. However, they can’t tell you whether it’s the paternal or the maternal copy.
Chromosome mapping is a puzzle in which you assign the segments of DNA you share with your DNA matches to the ancestors you inherited them from.
More help
- My free webinar An introduction to DNA Painter is available. If this leaves you cold, you could try Blaine Bettinger’s video introduction.
- There are many additional articles and videos available on this help page
It can definitely take a little while to get your head around the concepts. If you use Facebook, I would recommend joining the user group.
How do I make a match maternal or paternal in my chromosome map?
If you have a match that you want to make maternal or paternal, click on any of the match’s segments. This will bring up a popup.
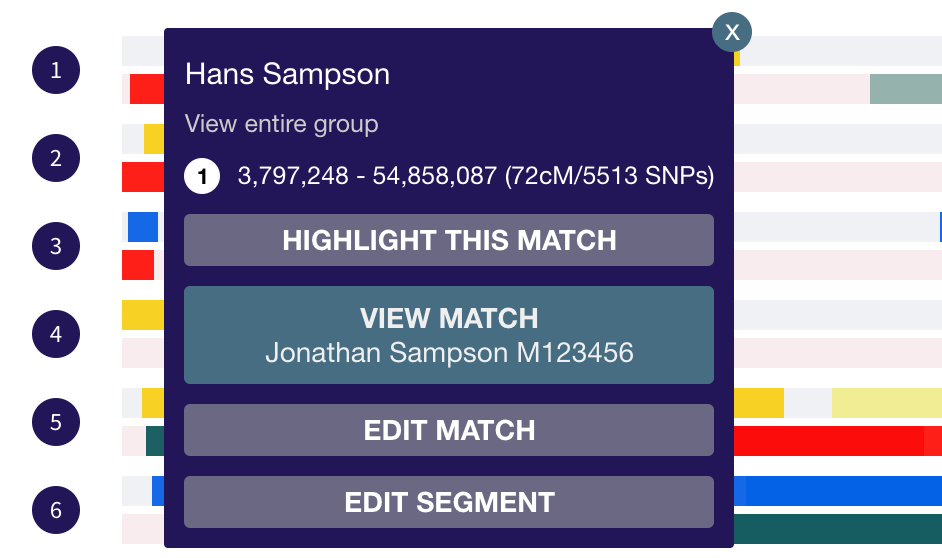

To edit the match:
- Click ‘edit match’
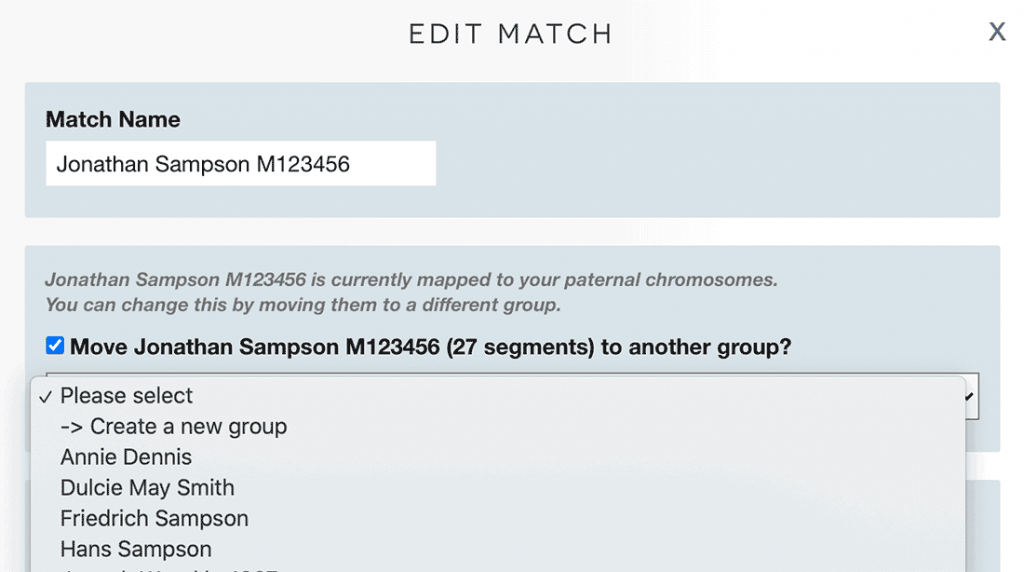
- Check the box that says ‘Move [match name] to another group’
- You can then select a different ancestor from the dropdown
- Or you can create a new group or ancestor that is specifically maternal or paternal
- Click ‘Save match’ when you’re finished
How do I assign a segment to a different ancestor?
You can edit segments individually just as you do matches. To edit a segment, click on it and then click ‘Edit segment’. You can assign that segment to a different group or ancestor or create a new one.
If people have overlapping segments on the same chromosome does that mean they are related?
Not necessarily. If two matches share segments with you that overlap visually, there are at least two possibilities:
- The overlapping portion is on the same copy of your chromosome, and they therefore share a common ancestor
- Each of them matches you on a different copy, and so the overlap does not imply that they are related
Assessing an overlapping match
In the example below, a new match has a segment that overlaps with a maternal aunt. This is great news. While it doesn’t immediately tell you how you’re related, it means that by comparing the two matches, you will be able to figure out which copy of your chromosome 1 the unidentified match is on.
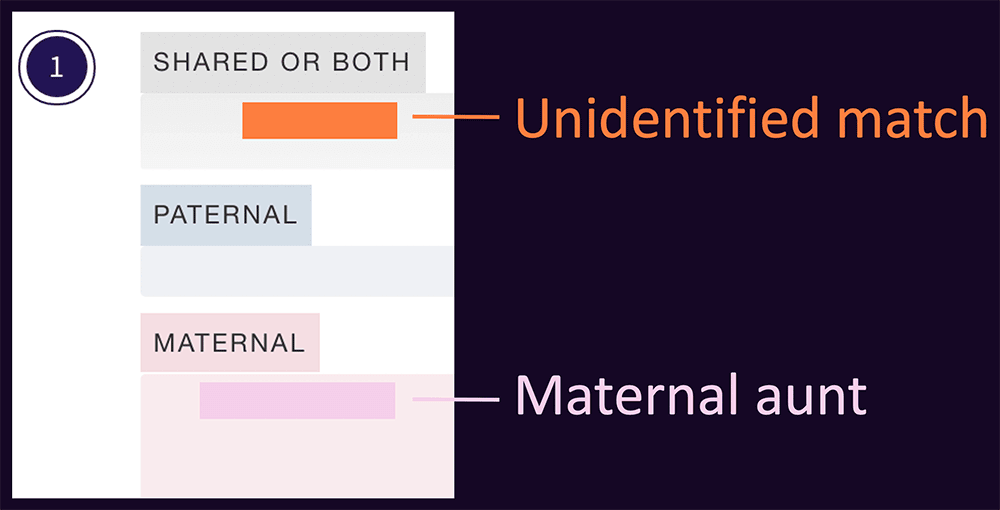
This works as follows
- If the unidentified person is a match to your aunt, then that segment is maternal
- If they don’t match your aunt, you can infer that it’s paternal
DNA Painter does not store any raw DNA, so to figure out whether or not two matches are related, you have to go to a testing platform:
- At Gedmatch and MyHeritage, you can compare two or more people directly
- At FamilyTreeDNA you can use the matrix tool to see if the people are a match to each other, but you can’t determine with certainty that they match on any specific segment.
Essentially, you’re looking for evidence that can tell you either
- ‘These people share the same piece of DNA with me’ or
- ‘These people both match me in this spot, but it’s not the same piece of DNA, so one is on the maternal copy of chromosome and the other is on my paternal copy’
You might find this article by Roberta Estes useful:
It’s also worth bearing in mind that the start and end points of matching segments are not as precise as the specific numbers imply. So, unless an overlap is significant (that is, more than just 1-5cM), it’s possible that there isn’t a true match at all.
I have match who has data at FamilyTreeDNA, Gedmatch and MyHeritage. Which segments should I use?
Data from any of these sites is fine and equally valid. In all cases you may need to cross-reference with other data if an apparent contradiction arises. Modern autosomal DNA tests are only testing certain SNPs within the genome, so any matching segments will never be absolutely precise.
When you’re mapping your chromosomes you can compare your matches to see if they make sense alongside each other. Small segments can sometimes be false positives, but when you’re mapping your chromosomes these will generally jump out at you.
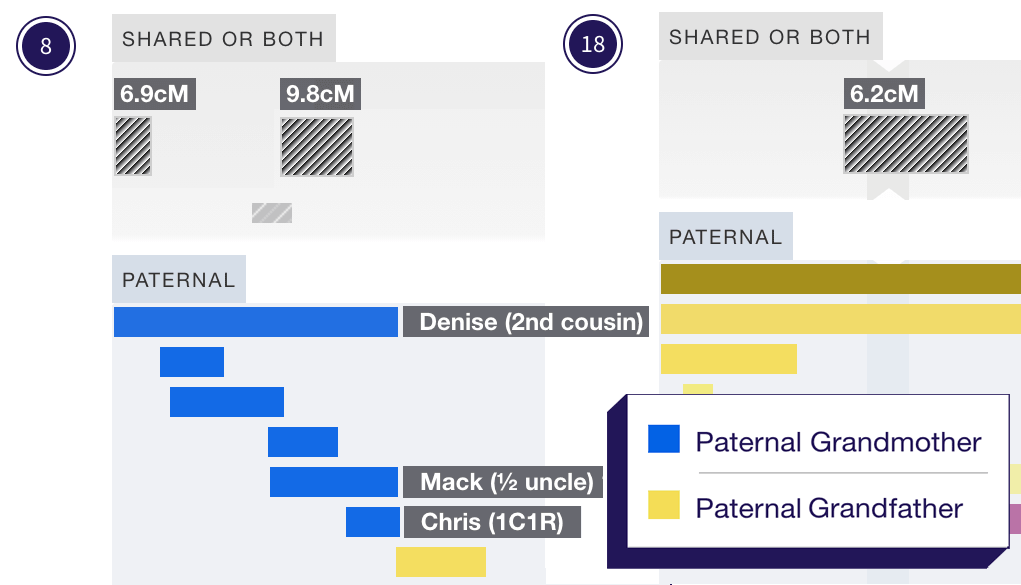
The smaller a matching segment, the more likely it is to be a coincidental match involving a weaving of DNA from both the maternal and paternal chromosomes. By default, DNA Painter will import only segments of 7cM or more, although you can adjust this threshold in the ‘Paint a match’ form.
Here are some facts to be aware of when working with segments from each different site:
- 23andme:
Segments down to 5cM are included. Smaller segments may be unreliableAs of late 2023, no segment data is available at 23andme. - FamilyTreeDNA:
Segments down to 1cM are included. This means that for every single match at FamilyTreeDNA, there will be many false segmentsAs of July 2021, only segments over 6cM are reported. You should still be wary of smaller segments. - Gedmatch: It may be necessary to check the ‘Prevent Hard Breaks’ option to prevent longer segments from being broken up
- MyHeritage: Segments down to 6cM are included. Sometimes segments can be erroneously extended beyond their true size.
How many matches on a chromosome would be needed to consider it a pile up?
There’s no absolute answer here. If I have 10 or more people and I can find no apparent link even after maxing out my research, I will consider it a likely pileup. But it could just be that the segment originates just beyond the genealogical timeframe. Many pileups will be more easily identifiable with dozens or even hundreds of matching segments in one place.
How can I back up my data?
From within any map, click on the settings cog above chromosome 1 and click ‘All segment data’. There’s a link in the top left to download your data in CSV format.
What are the Odds? and the Shared cM tool
What is the difference between WATO and WATO plus?
Overall, WATO plus has been overhauled with the aim of making it less confusing. Please read more here.
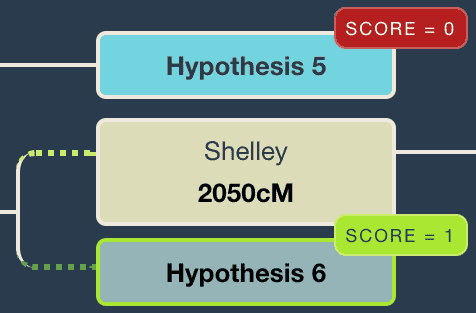
Why are the hypotheses sometimes different shades of blue?
WATO and WATO plus have a ‘Suggest hypotheses’ button. Any hypotheses that have been suggested by WATO are in a duller blue colour with a green background. In WATO plus the background is dotted.

The shared cM for a match differs across the testing sites. Which should I use in WATO/the Shared cM tool?
It’s unlikely to make a significant difference. Ancestry data benefits from having segments that are common in the population stripped out, while for some other sites it’s worth stripping out smaller and/or X segments. For further discussion please see Leah Larkin’s recent blog post.
Why are the scores different between v1 and v2 in WATO?
They are using a different set of probabilities. For more information, please see Leah Larkin’s summary.
Why does the target in WATO have to be born after 1900?
The target in the original WATO is the person who has taken a DNA test and whose matches you are using in the tree. For more background, please see the WATO FAQ. This is different in WATO plus.
Why is WATO saying it can’t suggest any hypotheses?
When you click ‘suggest hypotheses’, WATO will attempt to find positions in the tree that work for the cM numbers you’ve entered. When it can’t find any, this is usually because:
- Unknown additional relationships are inflating the cM numbers
- Or there is a mistake in the tree (for example, someone has been entered in the wrong generation

I always suggest adding a few hypotheses manually and clicking ‘View score calculation’ to see which individual relationships are invalidating them. This may help to point you towards the problem.
How do I indicate half relationships in WATO?
- Hover over any person in the tree and click ‘Edit half-relationships’
- Check a box against any siblings that this person has a half-relationship with
Did I miss anything? Please let me know!
Contact info: @dnapainter.bsky.social / jonny@dnapainter.com
