When a testing company provides the segments that you share with a DNA match, they will include an amount of centiMorgans (cMs) for each segment. But there may be times when you need to split a segment and assign each section to different ancestors. In this post I’ll introduce cM estimator, a new tool that can help you calculate the cMs in any segment or batch of segments.
Background
A centiMorgan is a unit of genetic distance based on the probability that there will be a recombination in specific areas of each chromosome. The more cMs there are in a segment, the more significant it is, but centiMorgans do not represent linear distance.
For example, a segment on chromosome 1 that starts at position 84,225,454 and ends at position 117,804,751 is about 36cM. By contrast, a segment with the same start and end position on chromosome 2 is only about 24cM1.
How this tool came to exist
Amy Williams is an associate professor of Computational Biology at Cornell University. Amy recently released two interesting tools for genealogists working with DNA alongside a useful blog. I highly recommend checking her site out at hapi-dna.org.
Amy and I met at Rootstech in Salt Lake City earlier this year. I mentioned the need for a tool to calculate centiMorgans based on chromosome, start and end position. A venerable existing tool does exist but is not straightforward to use and does not always function as expected.
I was aware that publicly available genetic map data exists, but was unclear on how to make use of this gigantic dataset. Amy was able to provide interpolated data and suggest a simple method for making these calculations2.
The cM estimator tool
The cM estimator is available at https://dnapainter.com/tools/cme.

Two ways to use it
You can either:
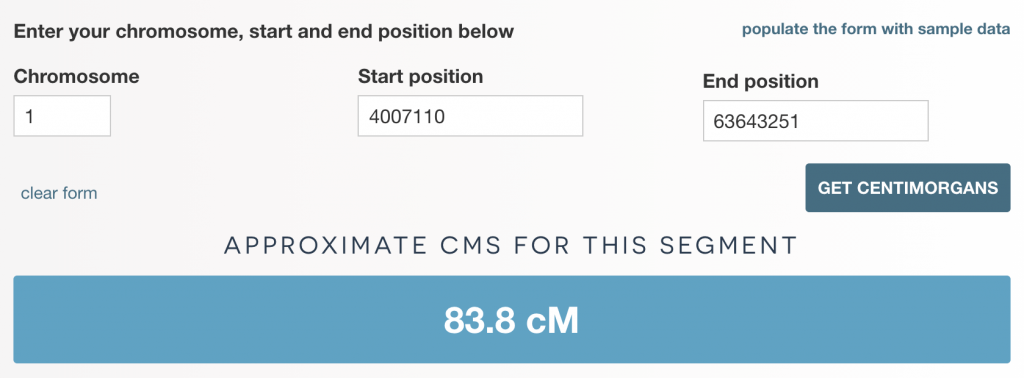
- enter the chromosome number, start and end position for an individual segment and click ‘GET CENTIMORGANS’
- OR copy and paste a batch of tab-delimited segments (chromosome start end) and click ‘GET CENTIMORGANS FOR A BATCH’
The results
The tool will then output the cMs for an individual segment, or output a set of tab-delimited rows with cMs for a batch of segments
Differences across sites
The eagle-eyed may notice that the cMs produced by the tool will not precisely match every other site. This could be for a number of reasons:
- The tool currently uses positions from the build 37 reference genome. This won’t match FamilyTreeDNA segments as they use build 36
- The method used, consulting a hapmap, seems to align closely with the cMs output by MyHeritage and 23andme
- The cMs are different from those by Gedmatch, who appear to be using a different method to calculate their figures
When would you use cM estimator?
I appreciate this may seem slightly obscure, but does actually have a few practical uses:
- If you’re comparing two matches where the segments overlap, this allows you to calculate how many cMs of DNA are in the overlapping part
- If you are attempting a visual phasing project, you will end up with segments where you just have the chromosome, start and end positions. This tool will allow you to confirm the cMs and relative significance of each segment.
cM estimator Integration with other tools
Perhaps more significantly, I’m also using these calculations in several upcoming tools. The first of these is the just-released Inferred segments generator.
Code
I’m releasing the code for the underlying API at https://github.com/dnapainter/apis
Footnotes
1. For a more technical explanation, I would recommend the blog post at Hapi-DNA: What is a centiMorgan?
2. You can read more on how the map was created in the article Minimal viable genetic maps
Contact info: @dnapainter.bsky.social / jonny@dnapainter.com
